Pannotator - Software
Some instructions
- Only the 'Annotation source 2' is not essential to job completion.
- Do not forget to submit your email address, if wrong email is provided we will not be able to mail you the result!
News
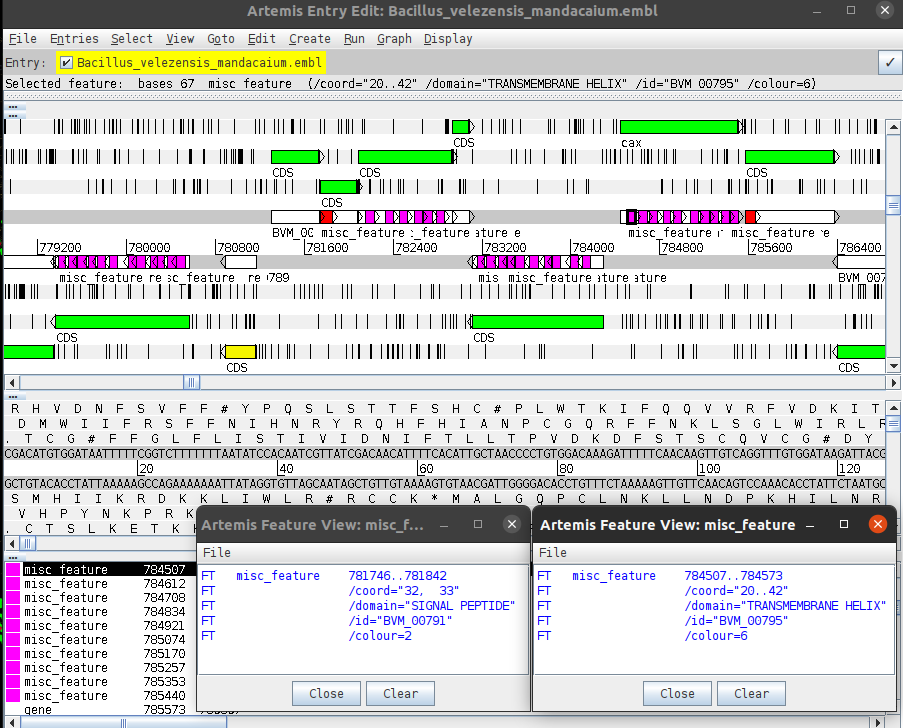
- This version works by calling ms-medpipe. The ms-medpipe starts the local subcellular predictions, but only PSE and SECRETED are tagged. Also, these proteins have the MED statistics calculated and normalized by the more significant score. The normalized MED score is assigned for its respective protein. Signal peptides and transmembrane domains are depicted at the DNA strand and linked to their host proteins. The absence of these motifs stands for cytoplasmatic proteins. Despite being quite useful for research, these features also should be manually removed before running SEQUIN. (2024)
- We adopted the pannotator to format and EMBL/GENBANK fitting the SEQUIN standards for genome submissions to NCBI. We are still adding the features 'colour' and 'annotation_source' to all annotated files for information. However, this data must be removed before running SEQUIN. (2023)
- rRNA and tRNA predictions updated. (2023)
- Google recaptcha (2022).
- Incorporated tRNA and mRNA predictions (2016).
- Pannotator optionally accepts a second curated file for annotations. The feature "annotation_source" keeps the record of the better match for each gene (2016).
- Pannotator is allowing the upload of a multifasta file for annotation. The service will independently annotate each DNA strand within the multifasta, and delivery all files in a compressed zip file (2015).
- Pannotator now uses the genbank reference file (EMBL, GBFF, or GBK) to train glimmer on how to predicted genes in the DNA strands, so no longer demands the upload of a CDS prediction file. Please, notice that the Uncompressed GBFF and GB file formats differ from each other. There is no DNA strand within a GB file composition which causes a processing error in the Pannotator software (2014). Unfortunately, we still have not a check for this failure, and the submitter one will receive nothing.
- Pannotator no longer demands specific file suffix names, just like FASTA, EMBL, GBFF or GBK. Uploaded files can have any acceptable nomenclature for the Linux OS files; however, files still need to be uploaded in the button designed for each type of file (2014).
- Gene's color tag depicting the alignment similarity among an annotated gene and it's annotation source: green (> 95%), yellow (< 95% and >= threshold%) and red (< trheshold%) (2013).
Bug fixes
- 1- Single annotation for each gene, no longer several superposed different annotations per gene.
- 2- Genome name entered in the moment of the submission with space delimiter causing delivery error.
Refer FAQs for file format details.